Annotation of the Covid-19 open research dataset for the scientific research community
In this blog find out how the SciBite team has responded to the tech community call to arms from The White House after they released an Open Research Dataset (CORD-19), with the hope to help uncover insights and answer high-priority scientific questions related to Covid-19.

In March 2020, in response to the global Coronavirus disease pandemic, a consortium including The White House released an Open Research Dataset (CORD-19) along with a call to arms to the tech community to help uncover insights and answer high-priority scientific questions related to Covid-19. This call asked for researchers around the world to apply or develop techniques to analyze and study a large collection of Covid-19 publication data which has been made open access to help the scientific community. The original datasets can be accessed on Kaggle.
Since then, SciBite has been continually working using our semantic data analytics software to produce biomedical ontology annotated versions of this data which we have been releasing to the public domain under a GPL license to offer a helping hand. Facilitated by our CTO, one of our Machine Learning experts along with help from our ontologies team, have used our software to produce 8 million sentence-level annotations on this dataset to help those trying to identify biomedical entities within the text.
Applying data science approaches, such as machine learning, can benefit from the normalization of biomedical concepts we have identified. Similarly, knowledge graph building can also exploit links between identified concepts in text, and extract them as relationships. The open annotation and publication of this data is our early contribution to this call to arms. A publication on this work is now available via bioRxiv.
Summary of Method
To identify relevant concepts, a focused set of ontologies was required with enrichments specific to concepts pertinent to COVID-19 research, as well as use our existing ontologies in broad areas of interest – drugs, genes, indications, phenotypes. These new Covid-19 specific ontologies developed were also released as vocabularies on our SciBite Labs website for use.
To create the annotations, we deployed TERMite, our named entity recognition engine (NER) to the text in the CORD-19 data (titles, abstracts, and body text where applicable). By injecting these results back into the original JSON as an additional set of objects, we aimed to prevent any compatibility issues that would have arisen for individuals and groups who had already begun work with the initial release of this data. In addition, to enable a more focused analysis, we also tabulated and transposed the annotated data to create a set of sentence cooccurrences to expedite sentence-focused relationship extraction.
Summary of results
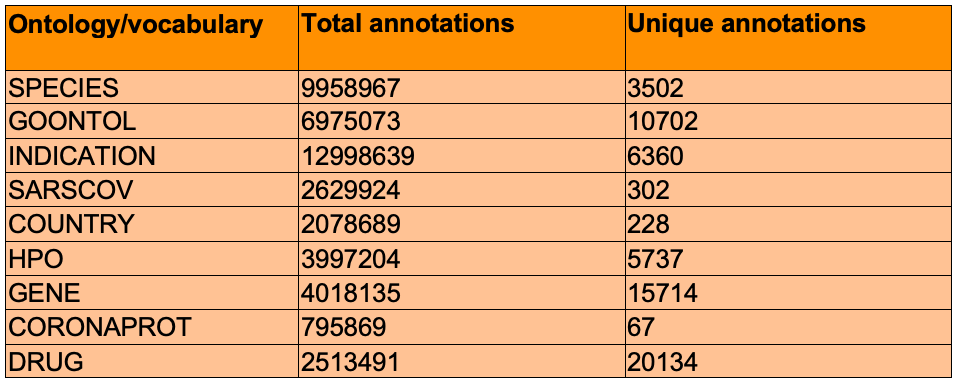
TERMite was able to identify and annotate over 45 million entities consisting of 62,746 unique ontology concepts. As can be observed in the table below, many of the drugs found both in March and more recently have been heavily investigated by researchers around the world for their use as either a treatment (e.g. Hydroxychloroquine, Ribavirin) or considering if they may play a role in poorer clinical outcome (e.g. Angiotensin).

The use of this data has already helped in developing knowledge graphs and data integration portals and it is our hope that it will be put to further use in the data-driven approaches used in 2020 and beyond.
To learn more take a look at the publication on this work which is now available via bioRxiv, or get in touch with the team if you’d like to find out more information.
Related articles
-
Drug repurposing, rare diseases and semantic analytics
In this blog we cover how to look potentially reduce the cost of and speed up the repurposing pipeline.
Read -
Rare disease collaboration networks
Disease Detective Part 1: In celebration of Rare Disease Day 28th Feb, we have a 3 part blog post looking into some of the challenges/analysis techniques involved in the research process.
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.