A report from the Biocuration 2023 Conference in Padua, Italy
 |
 |
The Biocuration Conference this year was held in the beautiful historic town of Padua in the Veneto region of Italy, renowned for its ancient University and picturesque old town. The stylish and relaxed atmosphere was the perfect place for catching up with old friends and establishing new connections and collaborations.

Article prepared with assistance from Rebecca Foulger, Paola Roncaglia and Jane Lomax
Before the conference proper, several workshops were held covering topics on standards for ontology mappings, aligning ontologies with Wikidata, and the intersection of biocuration and machine learning (ML). We learned how to correctly pronounce SSSOM* (as a spoiler, it’s not triple-S O M!), and discussions were held on the scope of the ontological mapping framework and use cases in the community. In the Machine Learning session, Jane Lomax (Head of Ontologies at SciBite) kicked off the panel discussion on the challenges of Machine Learning for Biocuration, with the panel providing their insights and advice on ML approaches.
The main conference opened strongly with a keynote from Paula Leitman of the Rio de Janeiro Botanical Garden, who described two bioinformatics systems to address the goals of the Global Strategy for Plant Conservation. The first is the Brazilian Flora and Funga System, which provides information on over 50,000 species of algae, fungi, and plants that occur in Brazil. The second is the Reflora Virtual Herbarium platform, a massive collaborative effort involving 76 herbaria around the world that aims to digitize images of Brazilian plants. An impressive achievement that will undoubtedly benefit the study of this rich biodiversity and its conservation.

SciBite sponsored the welcome drinks and nibbles
FAIR and share alike?
Still prominent in the conference schedule was the topic of FAIR (Findable, Accessible, Interoperable and Reusable) data. This was exemplified in the second keynote presentation from Marco Roos, one of the founders of the FAIR principles. It’s important to note again that the price of not adhering to FAIR is >10.2 billion Euros per year in Europe; this eye-watering statistic focused the collective mind of businesses and did a lot to kickstart the FAIR journey for many. Marco illustrated one example of a FAIR success story: Duchenne Muscular Dystrophy is one of the few rare diseases that now has a data registry, which acts as a central hub for patients and researchers. Marco showed a video of Lizanne Schreur, who was diagnosed with Duchenne. Lizanne described what FAIR and the easy access to information meant to her, not least the simple fact of knowing that she is not alone – and, importantly not the only woman who has this diagnosis – has an enormous impact. By reusing common infrastructure, Marco proposes that every rare disease will be able to easily set up a dedicated registry for the benefit of patients and researchers alike.
One thing of note from the meeting was that there is still some confusion between “FAIR” and “Open”, even in the Biocuration community. We must strive to make data adhere to the FAIR principles, but there is no obligation to make all data freely available. Think of companies that manufacture a product or institutes that use information from patient records; they want to be able to use their data within their system in the most efficient and economical way, i.e., FAIR, but they would not want to open up access to their proprietary information or sensitive patient data. Data should be “as open as possible, as closed as necessary.”
However, for resources that are open, there is also an obligation to keep them up-to-date and open in perpetuity. Charles Tapley Hoyt addressed this in his talk in the final conference session, which focused on how curated resources could be safeguarded from neglect, e.g., in the case of loss of funding, using the Bioregistry resource as an illustration. By providing not just open data, but open code, open infrastructure, and clear guidelines, a resource can be sustained, or revitalized, by community curators who would be rewarded with attribution and acknowledgment of their contributions.
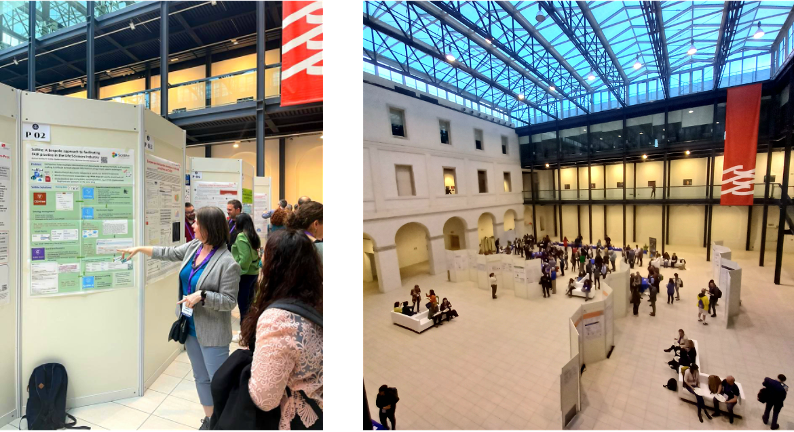

SciBite’s Lead Scientific Curator, Rachael Huntley in the Agorà of the Cultural Center Altinate San Gaetano, presenting a poster describing how SciBite helps customers make their data FAIR Biocuration 2023
Human in the loop
The shift in the types of technologies that biocurators make use of was evident from the very popular sessions on Artificial Intelligence (AI) and text mining approaches to curation. Where once there would have been talks on the automatic inference of functions, for example, there are now descriptions of machine learning and deep learning-based text mining.
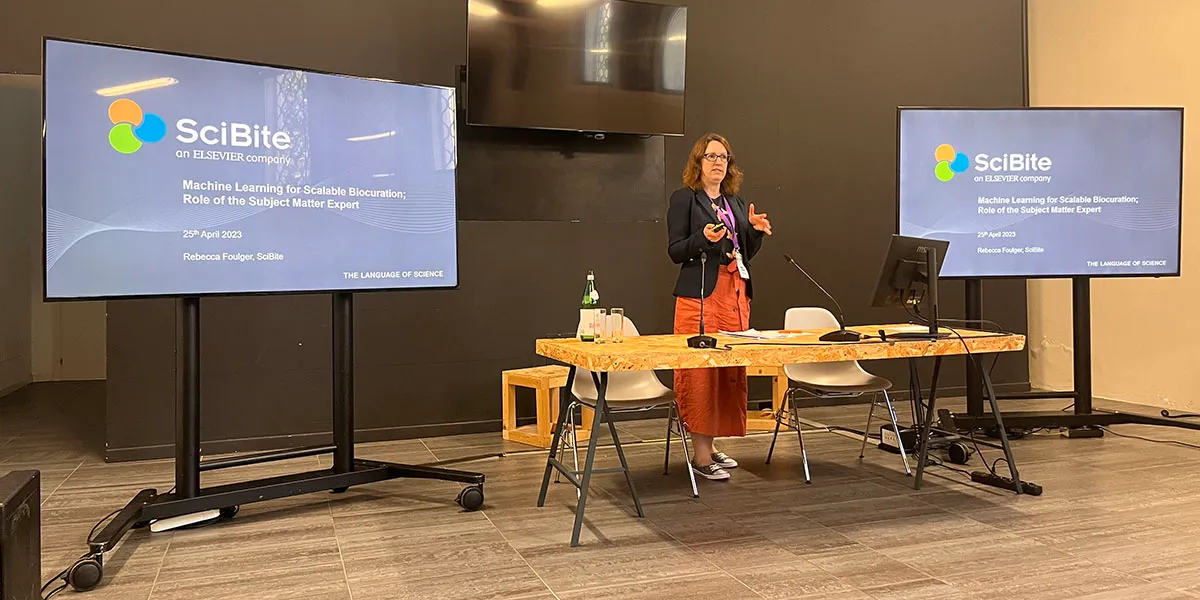
There were descriptions of using trained BERT models to extract variants and their impact on Alzheimer’s disease (C. Arighi “eMIND”), using text mining and ML approaches to identify entities and data types which are fed to community curators for verification (K. Van Auken “ACKnowledge”), ML classification of cell types (A. Niknejad), text mining-assisted extraction of protein-protein interactions (B. Gyori) and finally our own SciBite Senior Scientific Curator, Rebecca Foulger, describing how we incorporate ML to enrich our vocabularies with new terms or synonyms and to validate relationships between concepts.
The vital component, however, for these technologies to be a valuable asset to scaling up manual curation, efforts is the “human in the loop.” We are not at the point where ML or AI can reliably produce good quality, robust data. For instance, we still need gold-standard datasets to train models and expert biocurators to review and validate automatically produced data. Some biocurators were concerned that AI will replace them, but rather we need to put effort into workflows where biocurators and other Subject Matter Experts (SMEs) work alongside machine learning teams.
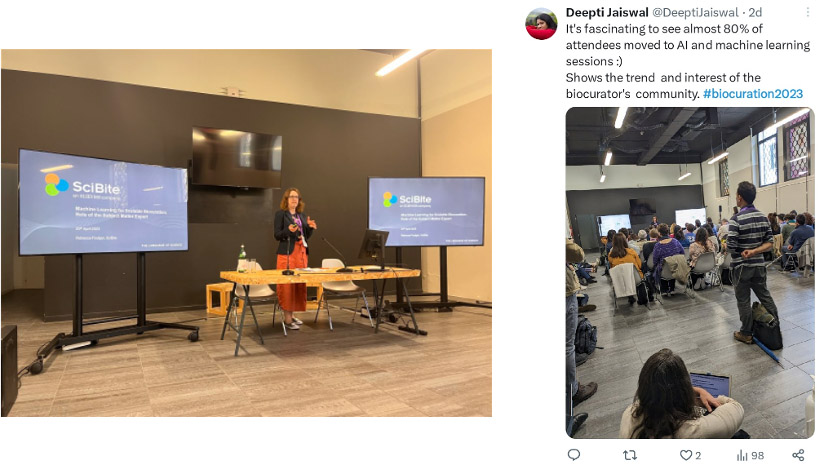
Standing room only for the AI and text mining session, in which SciBite’s Rebecca Foulger spoke on the role of the subject matter expert alongside machine learning
Finding your next role in biocuration
The conference also included a one-hour interactive Careers in Biocuration Panel/Workshop. During this time, general advice and suggested content for biocurators’ resumes/CVs was provided as well as more specific content, for those looking to join a particular organization. Attendees were also given up-to-date information on job application processes, including where to find job adverts and how CVs are pre-screened. The importance of adding keywords to CVs was highlighted, as there are likely HR systems that will prioritize CVs that match several keywords.
Throughout the workshop, we were also able to hear from individuals who have been involved in the hiring process, from a variety of sectors, including academia, industry, non-profits, and publishing. They provided us with insights into the hiring process in biocuration and biocuration-related fields, including their biggest DO’s and DON’Ts when writing a CV. For example, spelling mistakes from an applicant looking for a biocurator role, would, of course, be a big NO! Additionally, an interesting finding from this workshop, through audience participation, was that there are clear geographical differences when it comes to CV writing. For example, in the USA, when applying for biocurator roles in academia, there is no limit to the number of pages in your CV, and you are encouraged to list all your work/publications. This is a lot different to the UK, where there is usually a two-page limit! Finally, another great point from the audience was to apply the STAR method when writing a CV, as this helps an individual to list their accomplishments in a detailed but precise way.
In summary, this was an informative and engaging workshop, and all the points provided should be taken into consideration when an individual is looking for their next role in biocuration. Our only request is that the session should have lasted longer!

The conference dinner was held in the historical Caffè Pedrocchi
Summary
In all, the conference organizers did well to include a diverse range of topics outside the typical biocuration conference fare of human and model organism-centric curation. Attendees were from databases, academia, pharma companies, healthcare, commercial settings, and even a botanical garden. This provided an opportunity to make new connections and collaborations and to hopefully increase awareness of these important efforts. We even learned a new set of terminology (“synonym spaghetti” and “the crapome” being two of our favorites).
To conclude, as Jerven Bolleman from the Swiss Institute of Bioinformatics succinctly put it in his talk on FAIR knowledge graphs: 1) Structure all your information 2) Use ontologies 3) Identify everything.

Tweet from Jane Lomax ‘Crapome’ my favourite new expression from Yesterday @biocuration2023
*pronounced ‘sessom’ or S.S.S.O.M
Related articles
-
Creating a SciBite VOCab from a public ontology


Public ontologies are essential for applying FAIR principles to data but are not built for use in named entity recognition pipelines. At SciBite, we build on the public ontologies to create VOCabs optimized for NER. In this blog, discover how we create a SciBite VOCab from a Public Ontology.
Read -
The benefits of centralizing ontology management
Ontologies have become a key piece of infrastructure for organisations as they look to manage their metadata to improve the reusability and findability of their data. This is the first blog in our blog series 'Ontologies with SciBite'. Follow the blog series to learn how we've addressed the challenges associated with both consuming and developing ontologies.
Read
How could the SciBite semantic platform help you?
Get in touch with us to find out how we can transform your data
© Copyright © 2024 Elsevier Ltd., its licensors, and contributors. All rights are reserved, including those for text and data mining, AI training, and similar technologies.

